Cholera Outbreak 2022 in Karachi: A Report on Serotype and Antibiotic Susceptibility Pattern
By Qurat ul Ain Zahid, Nazia Khursheed, Fareeha Adnan, Adeel ZafarAffiliations
doi: 10.29271/jcpsp.2022.12.1613ABSTRACT
Objective: To identify the bacterial agent responsible for the surge of cases of acute diarrhoeal disease in Karachi, Pakistan, and assess the antimicrobial susceptibility pattern of the isolates.
Study Design: Observational study.
Place and Duration of Study: Department of Microbiology, Indus Hospital & Health Network, Karachi, Pakistan, from 1st March to 31st May 2022.
Methodology: Vibrio cholerae (V. Cholerae) isolates from stool samples received in the laboratory in the months of March, April, and May were identified and antibiotic susceptibility testing was performed by the Kirby-Bauer disk diffusion method. Serology was performed to identify the serotype causing a surge in cases of acute watery diarrhoea.
Results: A total of three-hundred and seventy-eight stool samples were received during the study period, out of which seventy-eight were positive for V. cholerae serogroup O1 serotype Ogawa. The isolate tested 100% sensitive for Ciprofloxacin, Tetracycline, Doxycycline, and Azithromycin, while 74% resistance was observed in Trimethoprim/Sulfamethoxazole and 25% resistance in Ampicillin.
Conclusion: V. cholerae serogroup O1 serotype Ogawa was the causative agent responsible for cholera outbreak in Karachi during the early summer of 2022. Ciprofloxacin, Tetracycline, Doxycycline, and Azithromycin are appropriate antimicrobials for treatment; whereas resistance to Trimethoprim/Sulfamethoxazole, and Ampicillin was higher.
Key Words: Vibrio cholerae, Outbreak, Acute diarrhoeal disease.
INTRODUCTION
Around 2 billion cases of acute diarrhoeal illness are reported across the globe annually. Acute watery diarrhoea is the second leading cause of death in children under the age of 5 years.1 Cholera is a water-borne acute diarrhoeal illness characterised by painless excretion of non-bloody watery stools resembling rice-water consistency, alkaline pH, and lack of pus cells.2-4 The causative agent of cholera is Vibrio cholerae (V. cholerae), which is a highly virulent, motile Gram-negative curved rod with the ability to release a toxin in the small intestine i.e., cholera toxin encoded by ctxAB genes.5 This toxin grants V. cholerae the potential to cause epidemic cholera which leads to severe dehydration, electrolyte imbalance, and shock. It can also result in a life-threatening condition called cholera gravis and even death if management is delayed.2,6,7
These characteristics distinguish cholera from other causes of acute diarrhoeal diseases and rule out major diarrhoeal pathogens such as Escherichia coli, Campylobacter spp., Salmonella spp., Shigella spp., and Yersinia enterocolitica.1 The factors contributing to the transmission include food and water contamination, poor sanitary systems, unhygienic practices, and lack of appropriate waste disposal.2,8-10
Cholera is a major health concern in developing countries, especially in Asian and African regions where it remains endemic.1,3,6,11,12 It is included in the list of national notifiable diseases.8 Pakistan gained the status of being endemic for cholera after 1988. The agricultural economy requires an extensive water distribution system which makes Pakistan prone to waterborne illnesses, particularly in metropolitan cities like Karachi due to overcrowding, unpredictable rainfalls, and poor sanitary system.2,11 Wastewater in Karachi has a high prevalence of toxigenic cholera which when combined with a poor sewerage system becomes a contributing factor in localised outbreaks of acute watery diarrhoea.2 In 1993 Karachi, Pakistan faced an epidemic of cholera caused by V. cholerae serogroup 0139, followed by serogroup O1 in 1994 with overlapping cases of both strains.5,7 Two more outbreaks were reported in Karachi, Pakistan, in July 2002 caused by V. cholerae O139 and in June 2003 by O1 serotype Ogawa.7
An upsurge in the cases of cholera was noticed in March, April, and May 2022. The purpose of this study was to identify the strain of V. cholerae responsible for the current outbreak, assess the antibiotic susceptibility profile of the isolate, and generate an antibiogram that will help clinicians in combating this disease and improving the chances of survival in susceptible individuals.
METHODOLOGY
Stool samples of all age groups, with complaints of watery diarrhoea visiting in the out-patient department (OPD), emergency department (ER) or admitted to the in-patient department (IPD), from 1st March to 31st May 2022 were included. Duplicate samples received from the same patient within 30 days were excluded. Samples were processed according to the standard operating procedure by following the American Society for Microbiology (ASM) guidelines.13 They were inoculated on Thiosulfate-citrate-bile-salt-sucrose agar (TCBS), Xylose-lysine deoxycholate agar (XLD), and MacConkey agar (MAC) plates. In addition, samples were also inoculated in alkaline peptone water (APW) for enhanced recovery. After overnight incubation at 35±2oC and ambient air for 24-48 hours, growth of V. cholerae was identified by performing Gram-stain of suspected colonies from TCBS agar. These colonies were streaked on nutrient agar and incubated at 35±2oC ambient air for 24 hours, biochemical tests such as oxidase and motility tests were performed. API 20 E strip (Biomérieux, Lyon, France) was used for definitive diagnosis. Serotyping of the isolated colonies was performed using polyvalent anti-serum (Ogawa and Inaba). Antimicrobial susceptibility testing of the isolate was performed on Mueller Hinton agar (MHA) using the Kirby-Bauer disk method. The panel of antibiotics tested includes Ampicillin (AMP), Tetracycline (TE), Ciprofloxacin (CIP), Trimethoprim/sulfamethoxazole (SXT), Doxycycline (DO), and Azithromycin (AZM). Zone sizes were measured according to the clinical and laboratory standards institute M45-A2 (CLSI).9 Zone size of AZM was measured according to the guidelines of the European Committee on Antimicrobial Susceptibility Testing (EUCAST) version 12.0.14
This observational study was carried out after acquiring approval from the Institutional Review Board (IRB) of the hospital (IHHN_IRB_2022_05_010). Data were retrieved from the electronic records of the hospital which included patient identifiers, demographics, and location/area of residence, culture results, and susceptibility pattern of antibiotics. The data was analysed using SPSS version 24.0 (IBM Corp., Armonk, NY). The distribution of data was tested from the Kolmogorov-Smirnov test. The descriptive statistics for age were reported as median (IQR). The frequencies and percentages were computed for all the categorical variables i.e. gender, screening results, and susceptibility pattern. The Chi-square test was applied to assess the association of organisms with age and gender. A p-value of <0.05 was taken as significant. A heat map for the distribution of cholera in different areas of Karachi was generated from the QGIS version 3.22.2.
RESULTS
A total of 378 samples were received for stool culture and 78 (20.63%) were found positive for V. cholerae. All the isolates were identified as serogroup O1 and serotype Ogawa. The median age of the sampled patients was 22 (IQR: 4-42) years. A clear female preponderance was seen in the positive cases and included 50 (64.10%) females and 28 (35.89%) males. A significant association was observed between the gender and culture results with a p-value of 0.034. Furthermore, the categorisation based on age groups revealed that 20 (25.6%) cases belonged to pediatric while 58 (74.3%) cases belonged to the adult population. The statistical association between the targeted groups is presented in Table I.
Table I: Association of the gender and age groups with cholera.
|
Variables |
Positive, n=78 |
Negative, n=300 |
p-value |
|
|
Gender |
Female |
50 (64.10) |
152 (50.66) |
0.034* |
|
Male |
28 (35.89) |
148 (49.33) |
||
|
Age Groups |
≤16 years |
20 (25.6) |
125 (41.6) |
0.010* |
|
≥17 years |
58(74.3) |
175 (58.3) |
||
|
* Statistically significant association. Chi-square test was applied to calculate p-values. |
||||
The data were also categorised based on the location of the patient. The highest number of cases were reported from Korangi 39 (50%) followed by Landhi 15 (19%). Moreover, 14 (18%) cases were reported from Bhittai colony, Mahmudabad, and Mehran town. The areawise distribution of the outbreak in the city is further depicted in the heat map presented in Figure 1.
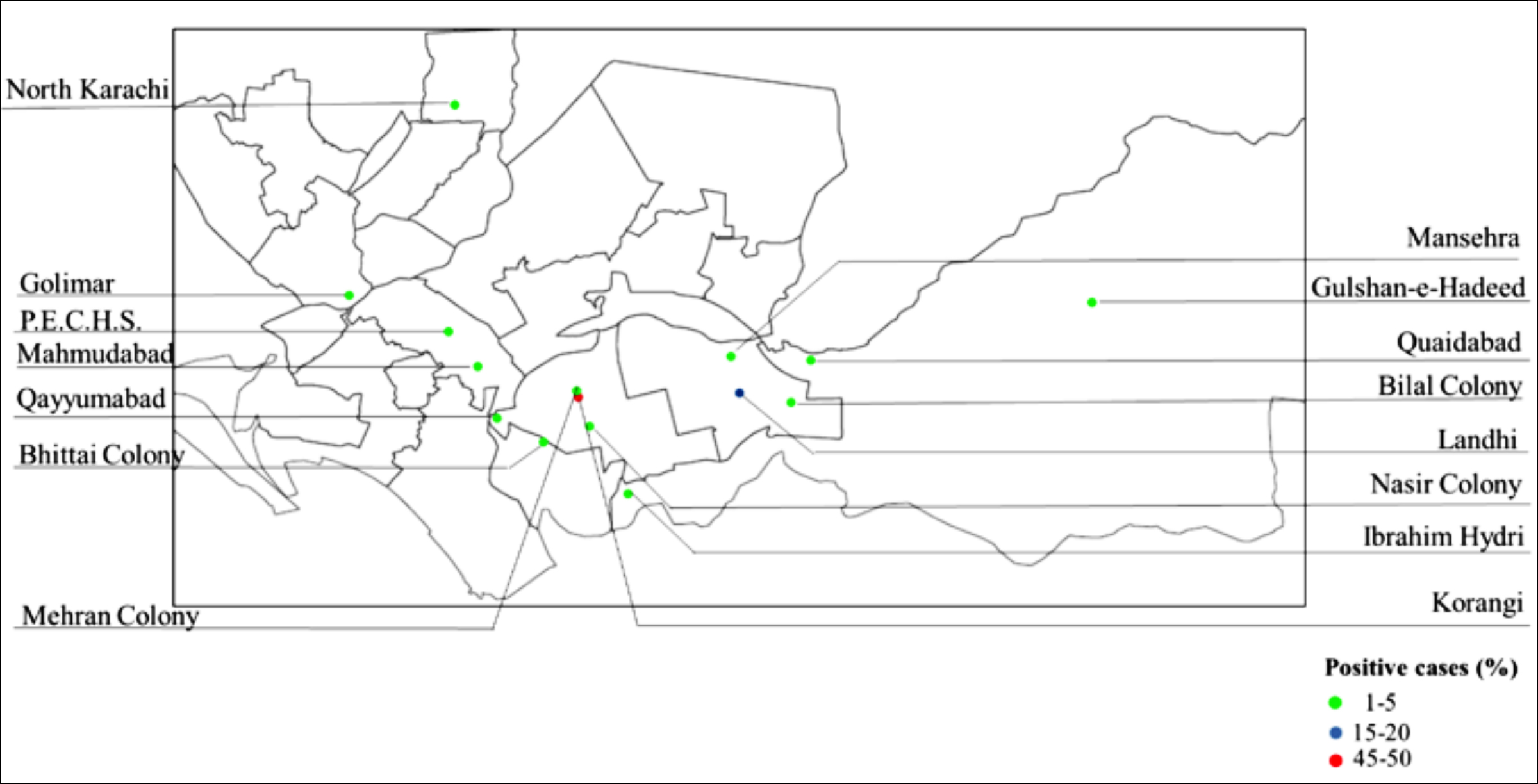 Figure 1: Areawise distribution of cholera in Karachi.
Figure 1: Areawise distribution of cholera in Karachi.
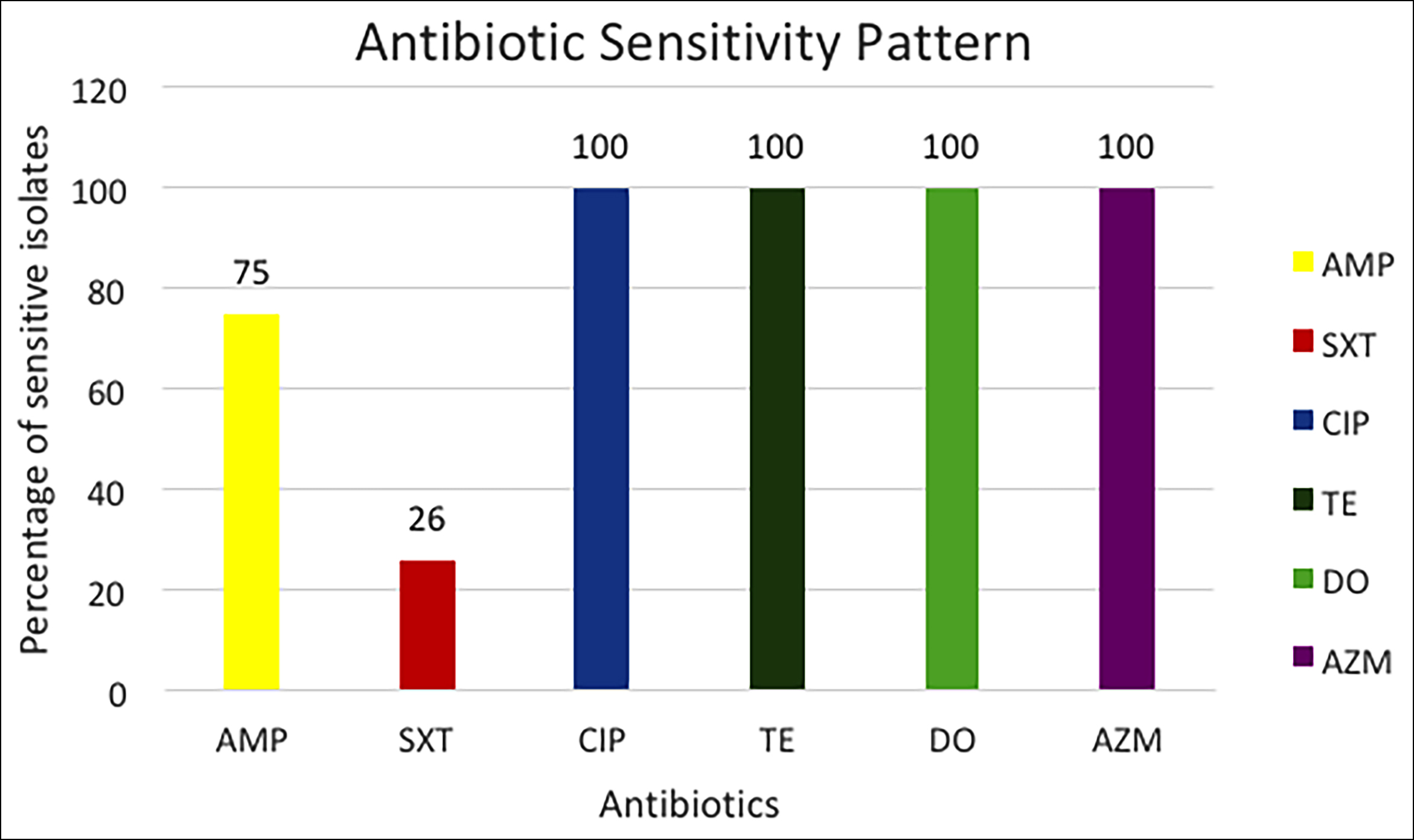 Figure 2: Antibiotic susceptibility pattern of isolated V. cholerae strains.
Figure 2: Antibiotic susceptibility pattern of isolated V. cholerae strains.
Among the isolated strains tested for first-line antibiotics, the highest resistance was observed in Trimethoprim/sulfamethoxazole (SXT) (74%) followed by Ampicillin (AMP) (25%) (Figure 2).
Whereas, all were found sensitive to Tetracycline (TE), Doxycycline (DO), Ciprofloxacin (CIP), and Azithromycin (AZM).
DISCUSSION
Outbreaks and epidemics are defined as an upward trend or sudden increase in the number of cases of a disease above the expected range for that population.11 In the last three months, the authors observed a sudden rise in the number of cases of cholera. Most of the patients were hospitalised due to severe dehydration and shock. The stool cultures were positive for V. cholerae O1 serotype Ogawa. There were an alarming number of cases from Korangi 39 (50%) followed by Landhi 15 (19%) and sporadic cases from other areas of Karachi. In the year 2021 there was not a single case reported of cholera between March and May, but this year there were 78 positive cases which were quite alarming and it was evident that an outbreak had occurred.
One of the positive findings of this study is the 100% sensitivity of isolated V. cholera to the four most commonly prescribed antibiotics i.e Ciprofloxacin, Tetracycline, Doxycycline, and Azithromycin.15 Sanford also recommends treatment of cholera with these antimicrobial agents.16 This current antibiogram will give confidence to general practitioners treating cholera patients. On the contrary, in a study conducted in Bangladesh in 2018, strain of V. cholerae was found to be resistant to tetracycline (14%), ciprofloxacin (1.5%), and azithromycin (1.5%).17 In another study conducted in Mozambique in 2015, 100% of the strains were resistant to ampicillin and tetracycline, 49% to azithromycin, and 95% to trimethoprim/sulfamethoxazole.18 By the year 2013, India reported 98.84% resistance to ampicillin, 89.53% to trimethoprim/sulfamethoxazole, 8.14% to tetracycline, and 1.16% to ciprofloxacin.19 In another cholera outbreak in Alborz Iran in 2011, a study was conducted and a resistance pattern of V. cholerae isolates was reported where 46% of the strains exhibited resistance to ampicillin, 95.4% to trimethoprim/sulfamethoxazole, 15.1% to tetracycline, and 4.2% to doxycycline.20 The present study exhibits sensitivities of ampicillin and trimethoprim/sulfamethoxazole in concordance with studies conducted in Bangladesh, India, and Mozambique.
The limitation of this study is being a single-centre study, including only those patients who visited the study centre and may not reflect the true disease burden in Karachi.
CONCLUSION
The serological testing of V. cholerae isolates identified serogroup O1 serotype Ogawa as the causative agent responsible for cholera outbreak in Karachi during the early summer of 2022. Ciprofloxacin, Tetracycline, Doxycycline, and Azithromycin are appropriate antimicrobials for treatment, whereas, the resistance of Trimethoprim/Sulfamethoxazole, and Ampicillin is rising. Cholera outbreak is alarming and highlights the need of improvement of drinking water disinfection and sanitation infrastructures including appropriate sewage disposal in Karachi, Pakistan.
ACKNOWLEDGEMENTS:
We are grateful to Nazia Parveen (Laboratory Supervisor) and the technical staff of the Department of Microbiology of Indus Hospital & Health network for handling the cases with great care and expertise. We are also thankful to Mamona Mushtaq (Research Associate) of Indus Hospital & Health Network Karachi for her assistance in statistical analysis.
COMPETING INTEREST:
The authors declared no competing interest.
AUTHORS’ CONTRIBUTION:
QZ: Contribution to concept, data collection, interpretation, and drafting the manuscript.
NK: Conceptualisation of the study, guidance in data collection, analysis and manuscript writing, and reviewing the content critically for final approval.
FA: Contribution to revising the initial draft, writing the manuscript, and reviewing it critically for final approval.
AZ: Contribution to revising the manuscript, data interpretation, statistical analysis, and reviewing it critically for final approval.
All authors took complete responsibility and accountability for all aspects of the work.
REFERENCES
- Ray S, Mukherjee T, Das A, Chattopadhyay B, Mukhopadhyay DK, Misra R. Cholera outbreak in Kamarhati municipality of West bengal: Secondary data analysis of a rapid epidemic response. J Compr Health 2021; 9(2):69-74.
- Zohra T, Ikram A, Salman M, Amir A, Saeed A, Ashraf Z, et al. Wastewater based environmental surveillance of toxigenic vibrio cholerae in Pakistan. PLoS One 2021; 16(9):e0257414. doi: 10.1371/journal.pone.0257414.
- Marin MA, Thompson CC, Freitas FS, Fonseca EL, Aboderin AO, Zailani SB, et al. Cholera outbreaks in nigeria are associated with multidrug resistant atypical El Tor and non-O1/non-O139 vibrio cholerae. PLoS Negl Trop Dis 2013; 7(2):e2049. doi: 10.1371/journal.pntd. 0002049.
- Awuor SO, Omwenga EO, Mariita RM, Daud, II. Cholera outbreak: Antibiofilm activity, profiling of antibiotic-resistant genes and virulence factors of toxigenic vibrio cholerae isolates reveals concerning traits. Access Microbiol 2022; 4(3):000324. doi: 10.1099/acmi.0. 000324.
- Sheikh A, Khan A, Malik T, Fisher-Hoch SP. Cholera in a developing megacity; Karachi, Pakistan. Epidemiol Infect 1997; 119(3):287-92. doi: 10.1017/s0950268897008212.
- Goswami S, Jha A, Sivan SP, Dambhare D, Gupta SS. Outbreak investigation of cholera outbreak in a slum area of urban Wardha, India: An interventional epidemiological study. J Family Med Prim Care 2019; 8(3):1112-6. doi: 10.4103/jfmpc.jfmpc_308_18.
- Siddiqui FJ, Bhutto NS, von Seidlein L, Khurram I, Rasool S, Ali M, et al. Consecutive outbreaks of vibrio cholerae O139 and V. cholerae O1 cholera in a fishing village near Karachi, Pakistan. Trans R Soc Trop Med Hyg 2006; 100(5):476-82. doi: 10.1016/j.trstmh.2005.07.019.
- Ministry of national health. Priority diseases for surveillance and response 2017 www.nih.org.pk/wp-content/uploads/ 2018/11/Notification-of-Priority-Diseases-in-Pakistan.pdf.
- CLSI. Methods for antimicrobial dilution and disc susceptibility testing of infrequently isolated or fastidious bacteria. 3rd ed. CLSI guidline M45. Wayne, PA: Clinical and laboratory standards institute; 2015.
- Usmani M, Brumfield KD, Jamal Y, Huq A, Colwell RR, Jutla A. A review of the environmental trigger and transmission components for prediction of cholera. Trop Med Infect Dis 2021; 6(3). doi: 10.3390/tropicalmed6030147.
- Mafi M, Goya MM, Hajia M. A five-year study on the epidemiological approaches to cholera in Iran. Caspian J Intern Med 2016; 7(3):162-7.
- Sodjinou VD, Talisuna A, Braka F, Barboza F, Alberti K, Fortin A, et al. The 2021 cholera outbreak in West Africa: Epidemiology and public health implications. Arch Clin Biomed Res 2022; 6(2):296-307.
- Leber AL. Clinical Microbiology Procedures Handbook. 4rth ed: American Society of Microbiology Press; 2016.
- The European Committee on Antimicrobial Susceptibility Testing. Routine and extended internal quality control for MIC determination and disk diffusion as recommended by EUCAST. Version 12.0, 2022.
- Das B, Verma J, Kumar P, Ghosh A, Ramamurthy T. Antibiotic resistance in Vibrio cholerae: Understanding the ecology of resistance genes and mechanisms. Vaccine 2020; 38 Suppl 1:A83-a92. doi: 10.1016/j.vaccine. 2019.06. 031.
- The Sanford Guide to antimicrobial therapy. 51st ed: Sperryvile, VA: Antimicrobial Therapy, Inc.; 2021.
- Parvin I, Shahunja KM, Khan SH, Alam T, Shahrin L, Ackhter MM, et al. Changing susceptibility pattern of vibrio cholerae O1 isolates to commonly used antibiotics in the largest diarrhoeal disease Hospital in Bangladesh during 2000-2018. Am J Trop Med Hyg 2020; 103(2):652-8. doi: 10. 4269/ajtmh.20-0058.
- Dengo-Baloi LC, Semá-Baltazar CA, Manhique LV, Chitio JE, Inguane DL, Langa JP. Antibiotics resistance in El Tor vibrio cholerae 01 isolated during cholera outbreaks in mozambique from 2012 to 2015. PLoS One 2017; 12(8):e0181496. doi: 10.1371/journal.pone.0181496.
- Pal BB, Nayak SR, Khuntia HK. Epidemiology and antibiogram profile of vibrio cholerae isolates between 2004-2013 from Odisha, India. Jpn J Infect Dis 2018; 71(2):99-103. doi: 10.7883/yoken.JJID.2017.193.
- Barati H, Moradi G, Rasouli MA, Mohammadi P. Epidemiologic and drug resistance pattern of vibrio cholerae O1 Biotype El Tor, serotype ogawa, in the 2011 cholera outbreak, in alborz province, Iran. Jundishapur J Microbiol 2015; 8(11):e23477. doi: 10.5812/jjm.23477.